Training Workshop for 2016 P30 CDAR Annual Meeting
Location: BST3, Room 3073, University of Pittsburgh
1:30 – 3:00 pm Core A Training (30 minute lecture and 1 hour demo & practice)
Training topics:
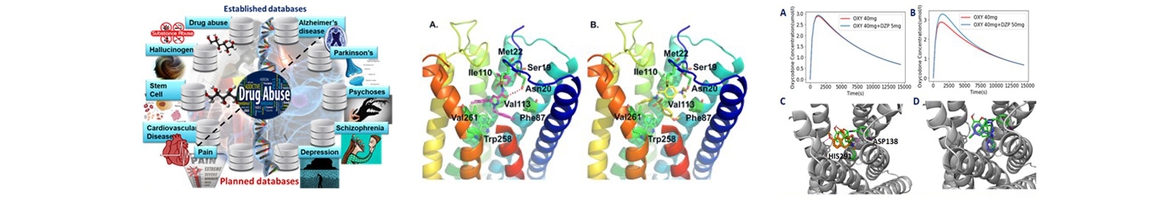
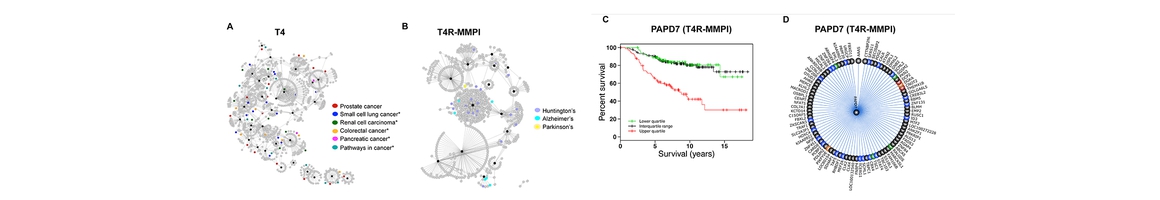
(1) Chemogenomics Database for Drug Abuse & Neuro-disorders: Genes, protein targets, pathways involved in a disease and small molecules that can directly interact with these key proteins with the potential to modulate the disease.
(2) TargetHunerMap: To predict the potential protein target(s) of a small molecule drug or a chemical compound through structure similarity or molecular docking.
(3) BBB prediction: To predict the blood-brain barrier (BBB) permeability of a small molecule based on its chemical features.
3:00 – 4:30 pm Core B Training (30 minute lecture and 1 hour demo & practice)
Training topics:
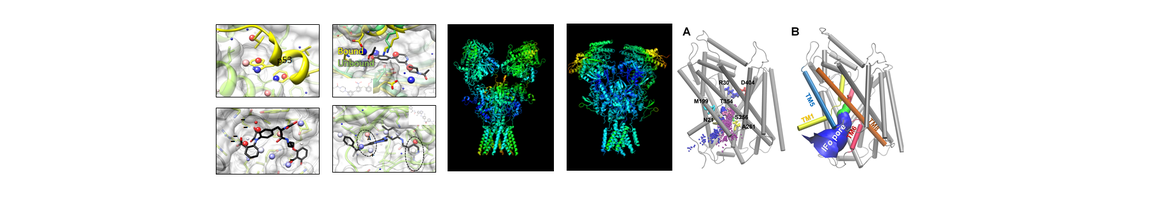
(1) ANM (Anisotropic network model): an elastic network based tool for analysis of dynamics of proteins and nucleic acids.
(2) ProDy: a free and open-source Python package for protein structural dynamics and sequence analysis.
(3)Druggability simulations: the use of DRUGUI, a VMD plugin, for performing MD simulations for druggability assessment, using probe molecules.
4:30 – 6:00 pm Core C Training (30 minute lecture and 1 hour demo & practice)
Training topic:
GenAMap: GenAMap software is a powerful platform for detection and easy visualization of structured association of genomic data and physical traits.