More than 180 publications and 3,300 citations to CDAR publications since 2014.
Year 2020
- Xing C, Zhuang Y, Xu T-H, Feng Z, Zhou XE, Chen M, Wang L, Meng X, Xue Y, Wang J, Heng Liu, McGuire TF, Zhao G, Melcher K, Zhang C*, Xu HE* and Xie X-Q*. Cryo-EM structure of human cannabinoid receptor CB2-Gi signaling complex. Cell, 2020 Jan 28. pii: S0092-8674(20)30054-4. doi: 10.1016/j.cell.2020.01.007. [Epub ahead of print]. PMID: 32004460
- Chen Y, Feng Z, Lin W, Wang Y, Wang S, Li C, Wang S, Chen M, Shan W* and Xie X-Q*. Effect of ginkgo biloba extract on the spatial learning and memory abilities in lithium-pilocarpine-induced status epilepticus rats: A combination of experimental study and computational systems pharmacology analysis. ACS Omega, 2020, in press. PMID: pending
- Journigan VB, Feng Z, Rahman S, Wang Y, Amin ARMR, Heffner CE, Bachtel N, Wang S, Gonzalez-Rodriguez S, Fernández-Carvajal A, Fernández-Ballester G, Hilton JK, Van Horn WD, Ferrer-Montiel A, Xie X-Q*, Rahman T*. Structure-based design of novel biphenyl amide antagonists of human transient receptor potential cation channel subfamily M member 8 channels (TRPM8) with potential implications in the treatment of sensory neuropathies. ACS Chem Neurosci 2020 Jan 9. [Epub ahead of print]. doi: 10.1021/acschemneuro.9b00404. PMID: 31850745
- Wang H, Vanyukov MM, Xing EP. & Wu W*. Discovering weaker genetic associations guided by known associations. BMC Medical Genetics, in press. PMID: pending
- Ji B, Liu S, He X, Man VH, Xie, X-Q, Wang J. Prediction of the binding affinities and selectivity for CB1 and CB2 ligands sing homology, molecular docking, molecular dynamics simulations, and MM-PBSA binding free energy calculations. ACS Chem Neurosci 2020, doi: 10.1021/acschemneuro.9b00696. [Epub ahead of print]. PMID: 32196303
- Ge S, Wang H, Alavi A, Xing EP, and Bar-Joseph Z. Supervised adversarial alignment of single-cell RNA-seq data. The 24th International Conference on Research in Computational Molecular Biology (RECOMB 2020). This is a refereed conference paper/proceeding.
- Zhai J, Liu S, Ji B, Zhang Y, Wang J. PBPK models of two CNS simulants, Amphetamine and methylphenidate, for clinical dosing regimen optimizations. Archives of Pharmaceuticals & Pharmacology Research. 2020, Article Number: 000545. doi: 10.33552/appr.2019.02.000545
- Li H, Doruker P, Hu G and Bahar I*. Modulation of toroidal proteins dynamics in favor of functional mechanisms upon ligand binding”, Biophysical Journal 2020, accepted. PMID: pending
- Li H, Pei F, Taylor DL, Bahar I. (2020) QuartataWeb: Integrated Chemical-protein-pathway Mapping for Polypharmacology and Chemogenomics. Bioinformatics 2020, in press.
- He X, Liu S, Lee TS, Ji B, Man V, York D, Wang J. Fast, accurate, and reliable protocols for routine calculations of protein-ligand binding affinities in drug
Year 2019
- Jing Y, Hu Z, Fan P, Xue Y, Wang L, Tarter RE, Kirisci L, Wang J, Vanyukov M, and Xie XQ. Analysis of substance use and its outcomes by machine learning I. Childhood evaluation of liability to substance use disorder. Drug Alcohol Depend. 2020 Jan 1; 206:107605. doi: 10.1016/j.drugalcdep.2019.107605. Epub 2019 Oct 22. PMID:31839402
- Hu Z, Jing Y, Xue Y, Fan P, Wang L, Vanyukov M, Kirisci L, Wang J, Tarter RE, and Xie XQ. Analysis of substance use and its outcomes by machine learning: II. Derivation and prediction of the trajectory of substance use severity. Drug Alcohol Depend. 2020 Jan 1;206:107604. doi: 10.1016/j.drugalcdep.2019.107604. Epub 2019 Oct 1. PMID:31615693
- Lee JY, Krieger JM, Li H, Bahar I. Pharmmaker: Pharmacophore modeling and hit identification based on druggability simulations. Protein Sci. 2020 Jan;29(1):76-86. doi: 10.1002/pro.3732. Epub 2019 Dec 4. PMID:31576621 PMCID:PMC6933858
- Zhang Y, Doruker P, Kaynak B, Zhang S, Krieger JM, Li H, Bahar I*. Intrinsic dynamics is evolutionarily optimized to enable allosteric behavior. Curr Opin Struct Biol. 2019 Nov 27;62:14-21. [Epub ahead of print]. doi: 10.1016/j.sbi.2019.11.002. PMID: 31785465
- Wang H, Wei Y, Cao M, Xu M, Wu W, & Xing EP. Deep inductive matrix completion for biomedical interaction prediction. IEEE International Conference on Bioinformatics and Biomedicine (IEEE BIBM 2019). This is a refereed conference paper/proceeding.
- Wu X, Mao Y, Wang H, Zeng X, Gao X, Xing EP, and Xu M., Regularized adversarial training (RAT) for robust cellular electron cryo tomograms classification. IEEE International Conference on Bioinformatics and Biomedicine (IEEE BIBM 2019). This is a refereed conference paper/proceeding.
- Wang Y, Guo H, Feng Z, Wang S, Wang Y, He Q, Li G, Lin W, Xie X-Q* and Lin Z*. PD-1-targeted discovery of peptide inhibitors by virtual screening, molecular dynamics simulation, and surface plasmon resonance. Molecules, 2019, 24(20), 3784; PMCID: PMC6833008; PMID: 31640203
- Zhang Y, Doruker P, Kaynak B, Zhang S, Krieger JM, Li H, Bahar I*. Intrinsic dynamics is evolutionarily optimized to enable allosteric behavior. Curr Opin Struct Biol. 2019 Nov 27;62:14-21. [Epub ahead of print]. doi: 10.1016/j.sbi.2019.11.002. PMID: 31785465
- Belovich AN, Aguilar JI, Mabry SJ, Cheng MH, Zanella D, Hamilton PJ, Stanislowski DJ, Shekar A, Foster JD, Bahar I, Matthies HJG, Galli A. A network of phosphatidylinositol (4,5)-bisphosphate (PIP2) binding sites on the dopamine transporter regulates amphetamine behavior in Drosophila Melanogaster. Mol. Psychiatry. 2019 Dec 3. [Epub ahead of print]. doi: 10.1038/s41380-019-0620-0. PMID:31796894
- Wang H, Lu C, Wu W & Xing EP. Graph-structured sparse mixed models for genetic association with confounding factors correction. IEEE International Conference on Bioinformatics and Biomedicine (IEEE BIBM 2019). This is a refereed conference paper/proceeding.
- Man VH*, He X, Ji B, Liu S, Xie XQ, and Wang J*. Molecular mechanism and kinetics of amyloid-β42 aggregate formation: A simulation study. ACS Chem Neurosci. 2019 Nov 20;10(11):4643-4658. Epub 2019 Nov 11. doi:10.1021/acschemneuro.9b00473. PMID:31660732
- Kim Y, Jun I, Shin DH, Yoon JG, Piao H, Jung J, Park HW, Cheng MH, Bahar I, Whitcomb DC, Lee MG. Regulation of CFTR Bicarbonate Channel Activity by WNK1: Implications for Pancreatitis and CFTR-Related Disorders. Cell Mol Gastroenterol Hepatol. 2020;9(1):79-103. doi: 10.1016/j.jcmgh.2019.09.003. Epub 2019 Sep 24. PMID:31561038
- Liu S, He X, Man VH, Ji B, Liu J*, and Wang J*. New application of in silico methods in identifying mechanisms of action and key components of anti-cancer herbal formulation YIV-906 (PHY906). Phys Chem Chem Phys 2019 Nov 14; 21(42):23501-23513. Epub 2019 Oct 16. doi: 10.1039/c9cp03803e. PMID:31617551
- Wang H, Yue T, Yang J, Wu W and Xing, EP*., 2019. Deep mixed model for marginal epistasis detection and population stratification correction in genome-wide association studies. BMC Bioinformatics, 20 (Suppl 23), pp.1-11, PMCID: PMC6933893.
- Cheng J, Wang S, Lin W, Wu N, Wang Y, Xie X-Q*, and Feng Z*. Computational System Pharmacology-Target Mapping for Fentanyl-laced Cocaine Overdose. ACS Chem. Neuro. 2019 Jul 15. doi: 10.1021/acschemneuro.9b00109. PMID: 31257858.
- Ma S, Attarwala I, Xie X-Q. SQSTM1/p62: a potential target for neurodegenerative disease. ACS Chem Neurosci 2019 Jan 18. doi: 10.1021/acschemneuro.8b00516.
- Wu N, Feng Z, He X, Kwon W, Wang J*, Xie X-Q*. Insight of Captagon Abuse by Chemogenomics Knowledgebase-guided Systems Pharmacology Target Mapping Analyses. Sci Rep 2019 Feb 19;9(1):2268: PMID: 30783122 PMCID: PMC6381188.
- Cheng MH, Bahar I. Monoamine transporters: structure, intrinsic dynamics and allosteric regulation, Nature Structural & Molecular Biology 2019; 26: 545–556.
- Taylor DL, Gough A, Schurdak ME, Vernetti L, Chennubhotla CS, Lefever D, Pei F, Faeder JR, Lezon TR, Stern AM, Bahar I. “Harnessing Human Microphysiology Systems as Key Experimental Models for Quantitative Systems Pharmacology” in Handbook of Experimental Pharmacology, 2019, p 1-41.
- Bian YM, He XB, Jing YK, Wang LR, Wang JM, Xie XQ. Computational systems pharmacology analysis of cannabidiol: a combination of chemogenomics-knowledgebase network analysis and integrated in silico modeling and simulation. Acta Pharmacologica Sinica. 2019 Mar;40(3):374. PMID: 30202014 PMCID: PMC6460368.
- Cheng MH, Ponzoni L, Sorkina T, Lee JY, Zhang S, Sorkin A, Bahar I. Trimerization of Dopamine Transporter Triggered by AIM-100 Binding: Molecular Mechanisms and Effect of Mutations. Neuropharmacology 2019 [Epub ahead of print]
- Zhou Z, Feng Z, Hu D, Yang P, Gur M, Bahar I, Cristofanilli M, Gradishar WJ, Xie XQ, Wan Y. A novel small-molecule antagonizes PRMT5-mediated KLF4 methylation for targeted therapy. EBioMedicine 2019; 19: 30312-30313. PMID: 31101597
- Pei F, Li H, Liu, B, Bahar I. Quantitative systems pharmacological analysis of drugs of abuse reveals the pleiotropy of their targets and the effector role of mTORC1. Front. in Pharm. 2019 [Epub ahead of print] PMID: 30906261
- Mikulska-Ruminska K, Shrivastava IH, Krieger JM, Zhang S, Li H, Bayir H, Wenzel SE, VanDemark AP, Kagan VE, Bahar I. Characterization of differential dynamics, specificity, and allostery of lipoxygenase family members. J Chem Inf Model. 2019 [Epub ahead of print] PMID: 30762363
- Wang Y, Lin W, Wu N, Wang S, Chen M, Lin Z, Xie X-Q*, and Zhi-wei Feng*. Structural insight into the serotonin (5-HT) receptor family by molecular docking, molecular dynamics simulation and systems pharmacology analysis. Acta Pharmacologica Sinica, 2019, in press.
- Zhang S, Li H, Krieger JM, Bahar I. Shared signature dynamics tempered by local fluctuations enables fold adaptability and specificity. Mol Biol & Evol 2019 [Epub ahead of print] PMID: 31028708
- Bian Y, Jing Y, Wang L, Ma S, Jun JJ, Xie XQ. Prediction of Orthosteric and Allosteric Regulations on Cannabinoid Receptors Using Supervised Machine Learning Classifiers. Mol Pharm. 2019 Jun 3;16(6):2605-2615. doi: 10.1021/acs.molpharmaceut.9b00182. Epub 2019 May 3. PMID: 31013097.
- Man, V.; Truong, P.; Li, M.; Wang, J.; Van-Oanh, N.-T.; Derreumaux, P.; Nguyen, P. Molecular Mechanism of the Cell Membrane Pore Formation Induced by Bubble Stable Cavitation, J. Phys. Chem. B, 2019, 123, 71-78
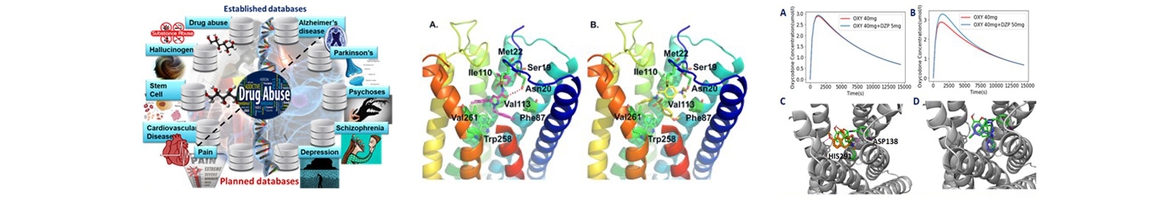
- Chen M, Jing Y, Wang L, Feng Z*, and Xie X-Q*. DAKB-GPCRs: An Integrated Computational Platform for Drug Abuse Related GPCRs. Journal of Chemical Information and Modeling, 2019, 59 (4), pp 1283–1289. DOI: 10.1021/acs.jcim.8b00623. PMID: 30835466.
- Fan P, Wang N, Wang L*, and Xie XQ*. Autophagy and apoptosis specific knowledgebases-guided systems pharmacology drug research. Current Cancer Drug Targets 2019 Feb 6;doi:10.2174/1568009619666190206122149. [Epub ahead of print]. PMID:30727895
- Wang H, Liu X, Tao Y, Ye W, Jin Q, Cohen JW, and Xing EP*. Automatic human-like mining and constructing reliable genetic association database with deep reinforcement learning, Proceedings of 24th Pacific Symposium on Biocomputing (PSB 2019). Publication is not a journal.
- Lee JY, Krieger J, Herguedas B, García-Nafría J, Dutta A, Shaikh SA, Greger IH, Bahar I. Druggability Simulations and X-ray Crystallography Reveal a Ligand-binding Site in the GluA3 AMPA Receptor N-terminal Domain. Structure 2019; 27: 241-252, PMID: 30528594
- Wang H, Wu Z, and Xing EP*. Removing confounding factors associated weights in deep neural networks improves the prediction accuracy for healthcare applications, Proceedings of 24th Pacific Symposium on Biocomputing (PSB 2019). Publication is not a journal.
- Ge, H. X.; Bian, Y. M.; He, H. X.; Xie, X. Q.; Wang, J. M., Significantly different effects of tetrahydroberberrubine enantiomers on dopamine D1/D2 receptors revealed by experimental study and integrated in silico simulation. J. Computer-Aided Molecular Design 2019, 33, 447-459. PMID: 30840169
- Wodak SJ, Paci E, Dokholyan NV, Berezovsky IN, Horovitz A, Li J, Hilser VJ, Bahar I, Karanicolas J, Stock G, Hamm P. Allostery in its many disguises: from theory to applications. Structure. 2019 Apr 2;27(4):566-578. doi: 10.1016/j.str.2019.01.003. Epub 2019 Feb 7. PMID: 30744993 PMCID: PMC6688844
- Wang L, Ma S, Hu Z, McGuire TF, Xie XQ. Chemogenomics Systems Pharmacology Mapping of Potential Drug Targets for Treatment of Traumatic Brain Injury. Journal of neurotrauma. 2019 Feb 1;36(4):565-75. PMID: 30014763 PMCID: PMC6354609 [Available on 2020-02-15] DOI: 10.1089/neu.2018.5757.
- Man VH, Li MS, Wang J, Derreumaux P, Nguyen PH. Interaction mechanism between the focused ultrasound and lipid membrane at the molecular level.J Chem Phys. 2019 Jun 7;150(21):215101. doi: 10.1063/1.5099008. PMID: 31176320
- Man VH, Li MS, Wang J, Derreumaux P, Nguyen PH. Nonequilibrium atomistic molecular dynamics simulation of tubular nanomotor propelled by bubble propulsion. The Journal of chemical physics. 2019 Jul 14;151(2):024103. PMID: 31301696 PMCID: PMC6624022
- Marchetti-Bowick M, Yu Y, Wu W, Xing EP. A penalized regression model for the joint estimation of eQTL associations and gene network structure. The Annals of Applied Statistics. 2019;13(1):248-70.
Year 2018
- Bertholomey ML, Stone K, Lam TT, Bang S, Wu W, Nairn AC, Taylor JR, Torregrossa MM*. Phosphoproteomic Analysis of the Amygdala Response to Adolescent Glucocorticoid Exposure Reveals G-Protein Coupled Receptor Kinase 2 as a Target for Reducing Motivation for Alcohol. Proteomes. 2018 Oct 12;6(4). pii: E41. doi: 10.3390/proteomes6040041. PMID:30322021 [Publication with FRP.]
- Kaya C, Cheng MH, Block ER, Bartol TM, Sejnowski TJ, Sorkin A, Faeder JR, Bahar I. Heterogeneities in Axonal Structure and Transporter Distribution Lower Dopamine Reuptake Efficiency. eNeuro. 2018:ENEURO. 0298-17.2017; PMCID: PMC5804147.
- Wang Y, Lin W, Wu N, He X, Wang J, Feng Z, Xie X-Q. An Insight into Paracetamol and Its Metabolites Using Molecular Docking and Molecular Dynamics Simulation. J Mol Model. 2018;24(9):243. PMID: 30121710.
- Wang Y, Lin W, Wu N, He X, Wang J, Feng Z, Xie X-Q. An Insight into Acetaminophen and Its Metabolites Using Molecular Docking and Molecular Dynamics Simulation. J Mol Model. 2018 Aug 18;24(9):243. PMID: 30121710 DOI: 10.1007/s00894-018-3790-9.
- Anthonymuthu T, Kenny E, Shrivastava I, Tyurina YY, Hier Z, Ting H-C, Dar H, Tyurin V, Nesterova A, Amoscato A, Mikulska-Ruminska K, Rosenbaum J, Mao G, Jinming Z, Conrad M, Kellum J, Wenzel S, VanDemark A, Bahar I, Kagan V, Bayir H (2018) Empowerment of 15-lipoxygenase catalytic competence in selective oxidation of membrane ETE-PE to ferroptotic death signals, HpETE-PE. J Am Chem Soc 2018, 140 (51), pp 17835-17839 PMID: 30525572.
- Wang K, Steer E, Otero PA, Bateman N, Cheng MH, Scott A, Wu C, Bahar I, Shih Y-T, Hsueh Y-P, Chu C (2018) PINK1 Interacts with VCP/p97 and Activates PKA to Promote NSFL1C/p47 Phosphorylation and Dendritic Arborization in Neurons. eNeuro 5 (6) ENEURO.0466-18.2018. PMID: 30783609
- Wu W*#, Bang S#, Bleecker ER, Castro M, Denlinger L, Erzurum SC, Fahy JV, Fitzpatrick AM, Gaston BM, Hastie AT, Israel E, Jarjour NN, Levy BD, Mauger DT, Meyers DA, Moore WC, Peters M, Phillips BR, Phipatanakul W, Sorkness RL, Wenzel SE* (# co-first authors). Multiview cluster analysis identifies variable corticosteroid response phenotypes in severe asthma”, Am J Resp Crit Care Med (accepted) [Publication with FRP.]
- Zhou L*, Zhou S, Yang P, Tian Y, Feng Z, Xie XQ*, Liu Y*. Targeted inhibition of the type 2 cannabinoid receptor is a novel approach to reduce renal fibrosis. Kidney Int. 2018 Oct;94(4):756-772. doi: 10.1016/j.kint.2018.05.023. Epub 2018 Aug 6. PMID:30093080.
- Lee JY, Krieger J, Herguedas B, García-Nafría J, Dutta A, Shaikh SA, Greger IH*, Bahar I*. Druggability simulations and X-ray crystallography reveal a ligand-binding site in the GluA3 AMPA receptor N-terminal domain. Structure. 2018 Nov 3. pii: S0969-2126(18)30378-2. doi: 10.1016/j.str.2018.10.017. [Epub ahead of print] PMID: 30528594
- Wang H, Lengerich BJ, Aragam B, Xing EP*. Precision Lasso: Accounting for Correlations and Linear Dependencies in High-Dimensional Genomic Data. Bioinformatics. 2018 Sep 1. doi: 10.1093/bioinformatics/bty750. PMID:30184048
- van Dijk L, Giladi M, Refaeli B, Hiller R, Cheng MH, Bahar I*, Khananshvili D*. Key residues controlling bidirectional ion movements in Na+/Ca2+ exchanger. Cell Calcium. 2018 Dec;76:10-22. PMID:30248574
- Wu X, Xie S, Wang L, Fan P, Ge S, Xie XQ, Wu W*. A computational strategy for finding novel targets and therapeutic compounds for opioid dependence. PLoS One. 2018 Nov 7;13(11):e0207027. doi: 10.1371/journal.pone.0207027. eCollection 2018. PMID:30403753 [Joint publication for Cores A and C.]
- Hu Z, Wang L, Ma S, Kirisci L, Feng Z, Xue Y, Klunk WE, Kamboh MI, Sweet RA, Becker J, Lv Q, Lopez OL*, Xie XQ*. Synergism of antihypertensives and cholinesterase inhibitors in Alzheimer’s disease. Alzheimers Dement (NY). 2018 Oct 14;4:542-555. PMID:30386819
- Krieger J, Lee JY, Greger IH, Bahar I. Activation and Desensitization of Ionotropic Glutamate Receptors by Selectively Triggering Pre-Existing Motions. Neurosci Lett. 2018:In press. doi: 10.1016/j.neulet.2018.02.050. PubMed PMID: 29481851; PMCID: PMC6107436.
- Wang, J; Ge, Y; Xie X-Q. Development and testing of druglike screening libraries. J. Chem. Inf. Model., Article ASAP (DOI: 10.1021/acs.jcim.8b00537).
- Man, V; He, X; Derreumaux, P; Ji, B; Xie X-Q; Nguyen, P; Wang, J. Effects of all-atom molecular mechanics force fields on Amyloid peptide assembly: the case of Aβ16-22 Dimer. J. Chem. Theor. Comput. (accepted) (Manuscript ID ct-2018-01107f.R1).
- Wang, J; Cieplak, P; Luo, R; Duan, Y. Development of Polarizable Gaussian Model for Molecular Mechanical Calculations I: Atomic Polarizability Parameterization to Reproduce Ab Initio Anisotropy. J. Chem. Theor. Comput. (accepted) (Manuscript ID ct-2018-006033.R2)
- He, X.; Man, V.; Ji, B.; Xie, X-Q; Wang, J. Calculate protein-ligand binding affinities with the extended linear interaction energy method: application on the Cathepsin S set in the D3R Grand Challenge 3. J. Comput.-Aided Mol. Des. 2018, Article ASAP (DOI: 10.1007/s10822-018-0162-6)
- Wang K, Steer E, Otero PA, Bateman N, Cheng MH, Scott A, Wu C, Bahar I, Shih Y-T, Hsueh Y-P, Chu CT* (2018) PINK1 interacts with VCP/p97 and activates PKA to promote NSFL1C/p47 phosphorylation and dendritic arborization in neurons. eNeuro, in press. PMID: pending
- Pei F, Li H, Liu B*, and Bahar I*. Quantitative systems pharmacological analysis of drugs of abuse reveals the pleiotropy of their targets and the effector role of mTORC1. Frontiers in Pharmacology, accepted with minor revision, doi: 10.1101/470922. PMID: pending
- Dominik, D.; Man, V.; Van-Oanh, N.-T.; Wang, J.; Kawasaki, T.; Derreumaux, P.; Nguyen, P. Breaking down cellulose fibrils with a mid-infrared laser, Cellulose, 2018, 25, 5553-5568.
- Xue Y, Feng ZW, Li XY, Hu ZH, Xu Q, Wang Z, Cheng JH, Shi HT, Wang QB, Wu HY, Xie X.-Q*. The efficacy and safety of cilostazol as an alternative to aspirin in Chinese patients with aspirin intolerance after coronary stent implantation: a combined clinical study and computational system pharmacology analysis. Acta Pharmacologica Sinica. 2018 Feb;39(2):205. PMID: 28933424 PMCID: PMC5800472
- Cheng MH, Kaya C, Bahar I. Quantitative Assessment of the Energetics of Dopamine Translocation by Human Dopamine Transporter. J Phys Chem B. 2018;122(21):5336-46. doi: 10.1021/acs.jpcb.7b10340. PubMed PMID: 29232131; PMCID: PMC5991911
- Cha-Molstad H, Lee SH, Kim JG, Sung KW, Hwang J, Shim SM, Ganipisetti S, McGuire T, Mook-Jung I, Ciechanover A, Xie XQ, Kim BY, Kwon YT. Regulation of autophagic proteolysis by the N-recognin SQSTM1/p62 of the N-end rule pathway. 2018 Jan 29:1-3. doi: 10.1080/15548627.2017.1415190. [Epub ahead of print] PMID:29261001
- Yoo YD, Mun SR, Ji CH, Sung KW, Kang KY, Heo AJ, Lee SH, An JY, Hwang J, Xie X-Q, Ciechanover A, Kim BY, Kwon YT. N-Terminal Arginylation Generates a Bimodal Degron That Modulates Autophagic Proteolysis. Proc Natl Acad Sci USA. 2018:201719110; PMCID: PMC5866579.
- Bian Y, Xie X-Q. Computational Fragment-Based Drug Design: Current Trends, Strategies, and Applications. AAPS J. 2018;20(3):59. PMID: 29633051 PMCID: PMC6618289 DOI: 10.1208/s12248-018-0216-7.
- Jing Y, Bian Y, Hu Z, Wang L, Xie X-Q. Deep Learning for Drug Design: An Artificial Intelligence Paradigm for Drug Discovery in the Big Data Era. AAPS J. 2018;20(3):58. PMID: 29603063 PMCID: PMC6608578
- Lengerich BJ, Aragam B, Xing EP. Personalized Regression Enables Sample-Specific Pan-Cancer Analysis. Bioinformatics (ISMB 2018). 2018;34(13):i178-86. doi: 10.1093/bioinformatics/bty250. PubMed PMID: 29949997; PMCID: PMC6022603.
- Sachan M, Xing EP. Self Training for Jointly Learning to Ask and Answer Questions. Proceedings of The 16th Annual Conference of the North American Chapter of the Association for Computational Linguistics (NAACL 2018); New Orleans, LA2018.
- Wang H, Liu X, Xiao Y, Xu M, Xing EP. Multiplex Confounding Factor Correction for Genomic Association Mapping with Squared Sparse Linear Mixed Model. Methods. 2018;145:33-40. Epub 2018/05/01. doi: 10.1016/j.ymeth.2018.04.020. PubMed PMID: 29705210; PMCID: PMC6108326.
- Xie P, Wu W, Zhu Y, Xing EP. Orthogonality-Promoting Distance Metric Learning: Convex Relaxation and Theoretical Analysis. The 35th International Conference on Machine Learning (ICML 2018)2018.
- Xie P, Xing EP. A neural architecture for automated icd coding. InProceedings of the 56th Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers) 2018 Jul (pp. 1066-1076).
- Dan C, Leqi L, Aragam B, Ravikumar PK, Xing EP. The sample complexity of semi-supervised learning with nonparametric mixture models. InAdvances in Neural Information Processing Systems 2018 (pp. 9321-9332).
- Xie P, Zhang H, Zhu Y, Xing EP. Nonoverlap-promoting variable selection. InInternational Conference on Machine Learning 2018 Jul 3 (pp. 5413-5422).
Year 2017
- Chen S, Feng Z, Wang Y, Ma S, Hu Z, Yang P, Chai Y, Xie, X.-Q*. Discovery of Novel Ligands for TNF-α and TNF Receptor-1 through Structure-Based Virtual Screening and Biological Assay. J Chem Inf Model. 2017 Apr 19. PMID: 28422491
- Bian Y, Feng Z, Yang P, Xie, X.-Q*. Integrated In Silico Fragment-Based Drug Design: Case Study with Allosteric Modulators on Metabotropic Glutamate Receptor 5. AAPS J. 2017 Jul;19(4):1235-1248.
- Cha-Molstad H, Yu JE, Feng Z, Lee SH, Kim JG, Yang P, Han B, Sung KW, Yoo YD, Hwang J, McGuire T, Shim SM, Song HD, Ganipisetti S, Wang N, Jang JM, Lee MJ, Kim SJ, Lee KH, Hong JT, Ciechanover A, Mook-Jung I, Kim KP, Xie, X.-Q*, Kwon YT*, Kim BY*. p62/SQSTM1/Sequestosome-1 is an N-recognin of the N-end rule pathway which modulates autophagosome biogenesis. Nature Communications 8, Article number: 102 (2017).
- Wang N, Wang L, Xie X.-Q. ProSelection: A Novel Algorithm to Select Proper Protein Structure Subsets for in Silico Target Identification and Drug Discovery Research. J Chem Inf Model. 2017 Nov 27;57(11):2686-2698. doi: 10.1021/acs.jcim.7b00277. Epub 2017 Oct 26. PMID:29016123
- Sauerwald N, Zhang S, Kingsford C, Bahar I. Chromosomal dynamics predicted by an elastic network model explains genome-wide accessibility and long-range couplings. Nucleic acids research. 2017 Apr 20;45(7):3663-73. PubMed PMID: 28334818; PubMed Central PMCID: PMC5397156.
- Cheng MH*, Torres-Salazar D*, Gonzalez-Suarez AD, Amara SG, Bahar I. (2017) Substrate transport and anion permeation proceed through distinct pathways in glutamate transporters.eLife 6: e25850.PMID: 28569666. (* equally contributed)
- Cheng MH, Garcia-Olivares J, Wasserman S, DiPietro J, Bahar I. (2017) Allosteric modulation of human dopamine transporter activity under conditions promoting its dimerization. J. Biol. Chem; in press.
- Lee J. Y., Feng Z., Xie X.-Q. , and Bahar I. (2017) Allosteric modulation of intact γ-secretase collective dynamics. Biophys. J. in revision.
- Li H, Sharma N, General IJ, Schreiber G, Bahar I. (2017) Dynamic Modulation of Binding Affinity as a Mechanism for Regulating Interferon Signaling. J Mol Biol; in press PMID:28648616.
- Li H, Chang Y-Y, Lee JY, Bahar I and Yang L-W. (2017) DynOmics: Dynamics of Structural Proteome and Beyond. Nucleic Acids Res. 45: W374-W380; PMID: 28472330.
- Ma S, Cheng MH, Guthrie DA, Newman AH, Bahar I, Sorkin A. (2017) Targeting of dopamine transporter to filopodia requires an outward-facing conformation of the transporter. Sci Rep. 7: 5399; PMID:28710426.
- Cheng, MH*, Garcia-Olivares, J*, Wasserman, S, DiPietro, J & Bahar, I. (2017) Allosteric modulation of human dopamine transporter activity under conditions promoting its dimerization. J. Biol. Chem 292:12471-12482; PMID: 28584050.
- Gur M, Cheng MH, Zomot E, Bahar I. (2017) Effect of dimerization on the dynamics of neurotransmitter: sodium symporters. J. Phys. Chem. B 121: 3657–3666. PMID: 28118712.
- Wang H, Aragam B, Xing EP. Variable Selection in Heterogeneous Datasets: A Truncated-rank Sparse Linear Mixed Model with Applications to Genome-wide Association Studies. KDD : proceedings. International Conference on Knowledge Discovery & Data Mining. bioRxiv. 2017 Jan 1:228106.
- Pei F, Li H, Henderson MJ, Titus SA, Jadhav A, Simeonov A, Cobanoglu MC, Mousavi SH, Shun T, McDermott L, Iyer P, Fioravanti M, Carlisle D, Friedlander RM, Bahar I, Taylor DL, Lezon TR, Stern AM, Schurdak ME. Connecting Neuronal Cell Protective Pathways and Drug Combinations in a Huntington’s Disease Model through the Application of Quantitative Systems Pharmacology. Sci Rep. 2017;7(1):17803; PMCID: PMC5736652.
- Xu, X. Chai, H. Muthakana, X. Liang, G. Yang, T. Zeev-Ben-Mordehai and E. P. Xing, “Deep learning based subdivision approach for large scale macromolecules structure recovery from electron cryo tomograms.” The Twenty-fifth International Conference on Intelligence Systems for Molecular Biology (ISMB 2017). Bioinformatics (2017) in press.
- Zhou Z, Luo A, Shrivastava I, He M, Huang Y, Bahar I, Liu Z, Wan Y. Regulation of XIAP turnover reveals a role for USP11 in promotion of tumorigenesis. EBioMedicine. 2017 Feb 1;15:48-61.PubMed PMID: 28040451; PubMed Central PMCID: PMC5233825.
- Xie X-Q, Wang L, Wang J, Xie Z, Yang P, Ouyang Q. Neuropathology of Drug Addictions and Substance Misuse. San Diego, CA. Elsevier Inc.;. Chapter 19, In silico chemogenomics knowledgebase and computational system neuropharmacology approach for cannabinoid drug research.
- Ponzoni L, Zhang S, Cheng MH, Bahar I. Shared Dynamics of LeuT Superfamily Members and Allosteric Differentiation by Structural Irregularities and Multimerization. Phil Trans R Soc B. 2018;373. doi: 10.1098/rstb.2017.0177. PubMed PMID: 29735731; PMCID: PMC5941172.
- Al-Shedivat M, Wilson AG, Saatchi Y, Hu Z, Xing EP. Learning Scalable Deep Kernels with Recurrent Structure. Journal of Machine Learning Research. 2017;18:1-37. PMID: 30662374 PMCID: PMC6334642
- Wenzel SE, Tyurina YY, Zhao J, Croix CMS, Dar HH, Mao G, Tyurin VA, Anthonymuthu TS, Kapralov AA, Amoscato AA, Mikulska-Ruminska K, Shrivastava IH, Kenny EM, Yang Q, Rosenbaum JC, Sparvero LJ, Emlet DR, Wen X, Minami Y, Qu F, Watkins SC, Holman TR, VanDemark AP, Kellum JA, Bahar I, Bayir H, Kagan VE. PEBP1 Wardens Ferroptosis by Enabling Lipoxygenase Generation of Lipid Death Signals. Cell. 2017;171(3):628-41. e26; PMCID: PMC5683852
- Neiswanger W, Xing EP. Post-Inference Prior Swapping. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017.
- Qin L, Zhang Z, Zhao H, Hu Z, Xing EP. Adversarial Connective-Exploiting Networks for Implicit Discourse Relation Classification. The 55th Annual Meeting of the Association for Computational Linguistics (ACL 2017); Vancouver, Canada.2017.
- Sachan M, Dubey A, Xing EP. From Textbooks to Knowledge: A Case Study in Harvesting Axiomatic Knowledge from Textbooks to Solve Geometry Problems. Proceedings of the 2017 Conference on Empirical Methods in Natural Language Processing (EMNLP 2017); Copenhagen, Denmark2017.
- Chang X, Yu Y, Yang Y, Xing EP. Semantic Pooling for Complex Event Analysis in Untrimmed Videos. IEEE Trans Pattern Anal Mach Intell. 2017;39(8):1617-32. Epub 2017/01/24. doi: 10.1109/tpami.2016.2608901. PubMed PMID: 28113653; PMCID: PMC5570670.
- Xie P, Deng Y, Zhou Y, Kumar A, Yu Y, Zou J, Xing EP. Analyzable Diversity-Promoting Latent Space Models. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017
- Xie P, Singh A, Xing EP. Uncorrelation and Evenness: A New Diversity-Promoting Regularizer. The 34th International Conference on Machine Learning (ICML 2017); Sydney, Australia2017.
- Xie P, Xing EP. A Constituent-Centric Neural Architecture for Reading Comprehension. The 55th Annual Meeting of the Association for Computational Linguistics (ACL 2017); Vancouver, Canada 2017.
- Yen IE, Huang X, Dai W, Ravikumar P, Dhillon I, Xing EP. Pd-Sparse : A Primal and Dual Sparse Approach to Extreme Multiclass and Multilabel Classification. The 23rd SIGKDD Conference on Knowledge Discovery and Data Mining (KDD2017); Halifax, Nova Scotia, Canada 2017.
- Zhang H, Zheng Z, Dai W, Q. H, Xing EP. Poseidon: An Efficient Communication Interface for Distributed Deep Learning on GPU Clusters. USENIX Annual Technical Conference; Santa Clara, CA, USA2017.
- Zhang K, Liu C, Zhang J, Xiong H, Xing EP, Ye J. Finding Structures of Large Matrices through Compression. The 23rd SIGKDD Conference on Knowledge Discovery and Data Mining (KDD2017); Halifax, Nova Scotia, Canada2017.
- Zhou Y, Yuan K, Yu Y, Ni X, Xie P, Xing EP, Xu S. Inference of Multiple-Wave Population Admixture by Modeling Decay of Linkage Disequilibrium with Polynomial Functions. Heredity (Edinb). 2017;118(5):503-10. Epub 2017/02/16. doi: 10.1038/hdy.2017.5. PubMed PMID: 28198814; PMCID: PMC5564381
- Lee S, Wang H, Xing EP. Backward genotype-transcript-phenotype association mapping. Methods. 2017 Oct 1;129:18-23. PMID: 28917724 DOI: 10.1016/j.ymeth.2017.09.004
- Xie P, Deng Y, Zhou Y, Kumar A, Yu Y, Zou J, Xing EP. Learning latent space models with angular constraints. InProceedings of the 34th International Conference on Machine Learning-Volume 70 2017 Aug 6 (pp. 3799-3810). JMLR. org
- Xie P, Salakhutdinov R, Mou L, Xing EP. Deep determinantal point process for large-scale multi-label classification. InProceedings of the IEEE International Conference on Computer Vision 2017 (pp. 473-482).
- Xie P, Poczos B, Xing EP. Near-Orthogonality Regularization in Kernel Methods. InUAI 2017 (Vol. 3, p. 6).
- Lee S, Görnitz N, Xing EP, Heckerman D, Lippert C. Ensembles of lasso screening rules. IEEE transactions on pattern analysis and machine intelligence. 2017 Nov 24;40(12):2841-52. PMID: 29989981
Year 2016
- Wang L, Xie X-Q. Cancer genomics: opportunities for medicinal chemistry? Future medicinal chemistry. 2016;8(4):357-359. doi:10.4155/fmc.16.1. PubMed PMID: 26976526; PubMed Central PMCID: PMC4847433.
- Hu J, Feng Z, Ma S, Zhang Y, Tong Q, Alqarni MH, Gou X, Xie X-Q. Difference and influence of inactive and active states of cannabinoid receptor subtype CB2: from conformation to drug discovery. Journal of chemical information and modeling. 2016 May 26;56(6):1152-63. PubMed PMID: 27186994; PubMed Central PMCID: PMC5395206.
- Feng Z, Pearce LV, Zhang Y, Xing C, Herold BK, Ma S, Hu Z, Turcios NA, Yang P, Tong Q, McCall AK. Multi-functional diarylurea small molecule inhibitors of TRPV1 with therapeutic potential for neuroinflammation. The AAPS journal. 2016 Jul 1;18(4):898-913. PubMed PMID: 27000851; PubMed Central PMCID: PMC5333490.
- Xu X, Ma S, Feng Z, Hu G, Wang L, Xie, X.-Q*. Chemogenomics knowledgebase and systems pharmacology for hallucinogen target identification-Salvinorin A as a case study. J Mol Graph Model. 2016 Nov;70:284-295. PMID: 27810775
- Zhang H, Ma S, Feng Z, Wang D, Li C, Cao Y, Chen X, Liu A, Zhu Z, Zhang J, Zhang G, Chai Y, Wang L, Xie, X.-Q*. Cardiovascular Disease Chemogenomics Knowledgebase-guided Target Identification and Drug Synergy Mechanism Study of an Herbal Formula. Sci Rep. 2016 Sep 28;6:33963. PMID: 27678063
- Zhang Y, Wang L, Feng Z, Cheng H, McGuire TF, Ding Y, Cheng T, Gao Y, Xie, X.-Q*. StemCellCKB: An Integrated Stem Cell-Specific Chemogenomics KnowledgeBase for Target Identification and Systems-Pharmacology Research. J Chem Inf Model. 2016 Oct 24;56(10):1995-2004. PMID: 27643925
- Hu J, Hu Z, Zhang Y, Gou X, Mu Y, Wang L, Xie, X.-Q*. Metal binding mediated conformational change of XPA protein: a potential cytotoxic mechanism of nickel in the nucleotide excision repair. J Mol Model. 2016 Jul;22(7):156. PMID: 27307058
- Stern AM, Schurdak ME, Bahar I, Berg JM, Taylor DL. A perspective on implementing a quantitative systems pharmacology platform for drug discovery and the advancement of personalized medicine. Journal of biomolecular screening. 2016 Jul;21(6):521-34. PubMed PMID: 26962875; PubMed Central PMCID: PMC4917453.
- Liu B, Liu Q, Yang L, Palaniappan SK, Bahar I, Thiagarajan PS, Ding JL. Innate immune memory and homeostasis may be conferred through crosstalk between the TLR3 and TLR7 pathways. Sci. Signal.. 2016 Jul 12;9(436):ra70-. PubMed PMID: 27405980; PubMed Central PMCID: PMC5087126.
- Li H, Chang YY, Yang LW, Bahar I (2016) iGNM 2.0: the Gaussian network model database for bimolecular structural dynamics Nucleic Acids Res 44: D415-422 PMID: 26582920.
- Kurkcuoglu Z, Bahar I, Doruker P (2016) ClustENM: ENM-based sampling of essential conformational space at full atomic resolution. J. Chem. Theory Comput. 12:4549-62; PMID: 27494295.
- Jun I*, Cheng MH*, Sim E*, Jung J, Suh BL, Kim Y, Son H, Park K, Kim CH, Yoon J-H, Whitcomb DC, Bahar I, Lee MG (2016) Pore Dilation Increases the Bicarbonate Permeability of CFTR, ANO1, and Glycine Receptor Anion Channels. Journal of Physiology 594: 2929-55; PMID:26663196.
- Lee, S, Lozano, A., Kambadur, P., and Xing EP “An Efficient Nonlinear Regression Approach for Genome-wide Detection of Marginal and Interacting Genetic Variations.” Journal of Computational Biology 2016, 23(5): 372-389.
- Lee S, Kong S, Xing EP. “A Network-driven Approach for Genome-wide Association Mapping.” Bioinformatics 2016, 32(12):i164-i173. doi: 10.1093/bioinformatics/btw270.
- Bang, S. and Wu, W. “Naive-Bayes Ensemble: A New Approach to Classifying Unlabeled Multi-Class Asthma Subjects.” The IEEE International Conference on Bioinformatics and Biomedicine (BIBM 2016).
- Marchetti-Bowick, J. Yin, J. Howrylak and E. P. Xing, A time-varying group sparse additive model for genome-wide association studies of dynamic complex traits, Bioinformatics 2016, 32 (19):btw347.
- Howrylak JA, Moll M, Weiss ST, Raby BA, Wu W, Xing EP. Gene Expression Profiling of Asthma Phenotypes Demonstrates Molecular Signatures of Atopy and Asthma Control. J Allergy Clin Immunol. 2016;137(5):1390-7.e6. Epub 2016/01/23. doi: 10.1016/j.jaci.2015.09.058. PubMed PMID: 26792209; PMCID: PMC4860055.
Year 2015 and before
- Feng, Z.; Pearce, L. V.; Xu, X.; Yang, X.; Yang, P.; Blumberg, P. M.; Xie, X.-Q. Structural Insight into Tetrameric hTRPV1 from Homology Modeling, Molecular Docking, Molecular Dynamics Simulation, Virtual Screening and Bioassay Validations. J. Chem. Inf. Model. 2015.
- Feng, Z.; Ma, S.; Hu, G.; Xie, X.-Q. Allosteric Binding Site and Activation Mechanism of Class C G-Protein Coupled Receptors: Metabotropic Glutamate Receptor Family. AAPS J. 2015, 1-17.
- Rentian Feng, Qin Tong, Zhaojun Xie, Haizi Cheng, Lirong Wang, Suzanne Lentzsch, G.David Roodman and Xiang-Qun Xie. Targeting cannabinoid receptor-2 pathway by phenylacetylamide suppresses the proliferation of human myeloma cells through mitotic dysregulation and cytoskeleton disruption. Molecular Carcinogenesis, DOI: 10.1002/mc.22251 16 JAN 2015
- Feng, Zhiwei, Stanton Kochanek, David Close, LiRong Wang, Ajay Srinivasan, Abdulrahman A. Almehizia, Prema Iyer, Xiang-Qun Xie, Paul A. Johnston, and Barry Gold. “Design and activity of AP endonuclease-1 inhibitors.” Journal of Chemical Biology (2015): 1-15.
- Yingdai Gao, Peng Yang, Hongmei Shen, Hui Yu, Xianmin Song, Liyan Zhang, Peng Zhang, Haizi Cheng, Zhaojun Xie, Sha Hao, Yahui Ding, Lirong Wang, Haibin Liu, Yanxin Li, Hui Cheng, Weimin Miao, Weiping Yuan, Youzhong Yuan, Tao Cheng, Xiang-Qun Xie* “Small-molecule inhibitors targeting INK4 protein p18 INK4C enhance ex vivo expansion of haematopoietic stem cells” Nature Communications. 02/2015; 6. DOI: 10.1038/ncomms7328
- Feng, Z., Ma, S., Hu, G., & Xie, X. Q. (2015). Allosteric binding site and activation mechanism of class C G-protein coupled receptors: metabotropic glutamate receptor family. The AAPS journal, 17(3), 737-753.
- Xiang-Qun Xie,* Lirong Wang, Haibin Liu, Qin Ouyang, Cheng Fang, and Weiwei Su.(2014) Chemogenomics knowledgebased polypharmacology analyses of drug abuse related G-protein coupled receptors and their ligands. Front Pharmacol. doi: 10.3389/fphar.2014.00003
- Ji, Kai-Long, Ping Zhang, Xiao-Nian Li, Juan Guo, Hua-Bin Hu, Chun-Fen Xiao, Xiang-Qun Xie, and You-Kai Xu. “Cytotoxic limonoids from Trichilia americana leaves.” Phytochemistry 118 (2015): 61-67.
- Myint, K. Z., & Xie, X. Q. (2015). Ligand Biological Activity Predictions Using Fingerprint-Based Artificial Neural Networks (FANN-QSAR). In Artificial Neural Networks (pp. 149-164). Springer New York.
- Teramachi, J.; Silbermann, R.; Yang, P.; Zhao, W.; Mohammad, K. S.; Guo, J.; Anderson, J. L.; Zhou, D.; Feng, R.; Myint, K. Z.; Maertz, N.; Beumer, J. H.; Eiseman, J. L.; Windle, J. J.; Xie, X. Q.; Roodman, G. D.; Kurihara, N., Blocking the ZZ domain of sequestosome1/p62 suppresses myeloma growth and osteoclast formation in vitro and induces dramatic bone formation in myeloma-bearing bones in vivo. Leukemia 2015.
- Silbermann, R., Zhou, D., Teramachi, J., Xie, X. Q., Roodman, G. D., & Kurihara, N. (2014). The p62-ZZ Domain Inhibitor XRK3F2 Alters Myeloma-Induced Suppression of Osteoblast Differentiation and Is Highly Cytotoxic to Myeloma Cells in Combination with Bortezomib. Blood, 124(21), 2083-2083.
- Zeng, D., Ouyang, Q., Cai, Z., Xie, X. Q., & Anderson, C. J. (2014). New cross-bridged cyclam derivative CB-TE1K1P, an improved bifunctional chelator for copper radionuclides. Chemical Communications, 50(1), 43-45.
- Zhang, S., Jia, N., Shao, P., Tong, Q., Xie, X. Q., & Bai, M. (2014). Target-Selective Phototherapy Using a Ligand-Based Photosensitizer for Type 2 Cannabinoid Receptor. Chemistry & biology, 21(3), 338-344.
- Wang, L., & Xie, X. Q. (2014). Computational target fishing: what should chemogenomics researchers expect for the future of in silico drug design and discovery?. Future medicinal chemistry, 6(3), 247-249.
- Liu, H., Wang, L., Su, W., & Xie, X. Q. (2014). Advances in recent patent and clinical trial drug development for Alzheimer’s disease. Pharmaceutical patent analyst, 3(4), 429-447.
- Feng, R., Milcarek, C. A., & Xie, X. Q. (2014). Antagonism of cannabinoid receptor 2 pathway suppresses IL-6-induced immunoglobulin IgM secretion.BMC Pharmacology and Toxicology, 15(1), 30.
- Feng, Z.; Alqarni, M. H.; Yang, P.; Tong, Q.; Chowdhury, A.; Wang, L.; Xie, X.-Q. Modeling, Molecular Dynamics Simulation and Mutation Validation for Structure of Cannabinoid Receptor 2 Based on Known Crystal Structures of GPCRs. J. Chem. Inf. Model. 2014, 54, 2483-2499.
- Qin Ouyang, Lirong Wang, Ying Mu, Xiang-Qun Xie, “Modeling skin sensitization potential of mechanistically hard-to-be-classified aniline and phenol compounds with quantum mechanistic properties”. BMC pharmacology & toxicology 12/2014; 15(1):76. DOI: 10.1186/2050-6511-15-76
- Lirong Wang, Chao Ma, Peter Wipf, Haibin Liu, Weiwei Su, Xiang-Qun Xie. TargetHunter: An In Silico Target Identification Tool for Predicting Therapeutic Potential of Small Organic Molecules Based on Chemogenomic Database. AAPS J. 2013 Apr; 15(2): 395–406. Published online 2013 Jan 5. doi: 10.1208/s12248-012-9449-z
- Cheng MH, Bahar I (2015) Molecular Mechanism of Dopamine Transport by Human Dopamine Transporter. Structure 23: 2171-81; PMID: 26106364.
- Gur M, Zomot E, Cheng MH, Bahar I (2015) Energy landscape of LeuT from molecular simulations. J Chem Phys 143: 243134; PMID: 26723619.
- Keskin O, Dyson HJ, Bahar I (2015) Biomolecular Systems Interactions, Dynamics, and Allostery. Reflections and New Directions. Biophys J 109: E01-2; PMID: 26377787.
- Krieger J, Bahar I, Greger IH (2015) Structure, Dynamics, and Allosteric Potential of Ionotropic Glutamate Receptor N-Terminal Domains. Biophys J 109: 1136-48; PMID: 26255587.
- Dutta A, Krieger J, Lee JY, Garcia-Nafria J, Greger IH, Bahar I (2015) Cooperative Dynamics of Intact AMPA and NMDA Glutamate Receptors: Similarities and Subfamily-Specific Differences. Structure 23; 1692-1704; PMID: 26256538.
- Haliloglu T, Bahar I (2015) Adaptability of protein structures to enable functional interactions and evolutionary implications. Current Opinion in Structural Biology 35: 17-23; PMID: 26254902.
- Bahar I, Cheng MH, Lee JY, Kaya C, Zhang S (2015) Structure-Encoded Global Motions and Their Role in Mediating Protein-Substrate Interactions. Biophys J 109: 1101-9; PMID: 26143655.
- Cheng MH, Block E, Hu F, Cobanoglu MC, Sorkin A, Bahar I (2015) Insights into the modulation of dopamine transporter function by amphetamine, orphenadrine, and cocaine binding; Front Neurol 6: 134; PMID: 26106364.
- Eyal E, Lum G, Bahar I (2015) The Anisotropic Network Model web server at 2015 (ANM 2.0). Bioinformatics 2015: 1-3; PMID: 25568280.
- Zomot E, Gur M, Bahar I. Microseconds Simulations Reveal a New Sodium-Binding Site and the Mechanism of Sodium-Coupled Substrate Uptake by LeuT. J Biol Chem. 2015;290(1):544-55. doi: 10.1074/jbc.M114.617555. PubMed PMID: 25381247; PMCID: PMC4281755.
- Cheng MH, Bahar I (2014) Complete Mapping of Substrate Translocation Highlights the Role of LeuT N-terminal Segment in Regulating Transport Cycle. PLoS Comput Biol 10: e1003879; PMID: 25299050.
- Cobanoglu MC, Oltvai ZN, Taylor DL, Bahar I (2014) BalestraWeb: Efficient, online evaluation of drug-target interactions. Bioinformatics 31:131-3; PMID: 25192741.
- Bahar I (2014) Coupling between Neurotransmitter Translocation and Protonation State of a Titratable Residue during Na(+)-Coupled Transport. Biophys J 106: 2547-8 PMID: 24940770.
- Bakan A,* Dutta A,* Mao W, Liu Y, Chennubhotla C, Lezon TR, Bahar I (2014) Evol and ProDy for Bridging Protein Sequence Evolution and Structural Dynamics. Bioinformatics 30: 2681-3; PMID: 24849577.
- Das A, Gur M,* Cheng MH,* Jo S, Bahar I, Roux B (2014) Exploring the conformational transitions of biomolecular systems using a simple two-state anisotropic network model; PLoS Comput Biol 10: e1003521 PMID: 24699246.
- S. Lee, A. Lozano, P. Kambadur, and E. P. Xing, An Efficient Nonlinear Regression Approach for Genome-Wide Detection of Marginal and Interacting Genetic Variations, to appear in the 19th International Conference on Research in Computational Molecular Biology (RECOMB 2015)
- S. Lee, J. K. Kim, X. Zheng, Q. Ho, G. A. Gibson, and E. P. Xing, On Model Parallelization and Scheduling Strategies for Distributed Machine Learning, Advances in Neural Information Processing Systems 28 (eds. Corinna Cortes and Neil Lawrence), MIT Press, 2014. (NIPS 2014).
- Wu, W.*, Bleecker, E., Moore, W., Busse, W. W., Jarjour, N., Castro, M., Chung, K. F., Calhoun, W. J., Erzurum, S., Gaston, B., Israel, E., Curran-Everett, D., Wenzel, S. E.* (2014) “Unsupervised phenotyping of Severe Asthma Research Program participants using expanded lung data.” Journal of Allergy and Clinical Immunology 133(5):1280-1288. NIHMS562128. (PMCID: PMC4038417).
- E. P. Xing, R. Curtis, G. Schoenherr, S. Lee, J. Yin, K. Puniyani, W. Wu, P. Kinnaird, GWAS in a Box: Statistical and Visual Analytics of Structured Associations via GenAMap, PLoS One, Volume 9, Issue 6, e97524, 2014.
- Kolar M, Liu H, Xing EP. Graph Estimation from Multi-Attribute Data. Journal of Machine Learning Research. 2014;15(May):1713-50. PubMed PMID: 25620892; PMCID: PMC4303188.
- Feng Z, Hu G, Ma S, Xie X-Q. Computational Advances for the Development of Allosteric Modulators and Bitopic Ligands in G Protein-Coupled Receptors. AAPS J. 2015;17(5):1080-95; PMCID: PMC4540734.
- Chang X, Yang Y, Xing E, Yu Y. Complex event detection using semantic saliency and nearly-isotonic SVM. InInternational Conference on Machine Learning 2015 Jun 1 (pp. 1348-1357).
- Alqarni M, Myint KZ, Tong Q, Yang P, Bartlow P, Wang L, Feng R, Xie X-Q. Examining the Critical Roles of Human CB2 Receptor Residues Valine 3.32 (113) and Leucine 5.41 (192) in Ligand Recognition and Downstream Signaling Activities. Biochem Biophys Res Commun. 2014;452(3):334-9; PMCID: PMC4465889.
- Zheng X, Yu Y, Xing EP. Linear time samplers for supervised topic models using compositional proposals. InProceedings of the 21th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 2015 Aug 10 (pp. 1523-1532). ACM.
Manuscripts submitted